This article was published in Scientific American’s former blog network and reflects the views of the author, not necessarily those of Scientific American
.png?w=590)
Perhaps you saw this graphic on the front page of The New York Times last week, leading into Amy Harmon's article about scientists from a variety of labs banding together in the fight against the Zika virus. The researchers' shared goal; sequence the genome of the virus' mosquito vector, Aedes aegypti, in the hope that a more complete knowledge of the insect’s genetic makeup will lead to ideas on how to prevent it from transmitting the virus that causes disease in humans. (The last major—although incomplete—sequencing effort was published in 2007).
The New York Times caption (as it appears online) states that you're looking at "A visualization of the recently sequenced Aedes aegypti genome. Each of the 3,752 colored lines is a fragment of its three chromosomes..."
On supporting science journalism
If you're enjoying this article, consider supporting our award-winning journalism by subscribing. By purchasing a subscription you are helping to ensure the future of impactful stories about the discoveries and ideas shaping our world today.
But what does that mean? How do you read the graphic, and how was it built? To find out, I reached out to Mark Kunitomi, author of the chart and postdoctoral fellow in the Andino Lab at University of California, San Francisco.
The genome sequence data for this chart was produced by the Andino lab in collaboration with Pacific BioSciences. As noted in Harmon’s article, other sequencing approaches are also currently being pursued, to refine the map further. (To learn more about a variety genome-reading technologies, see "Genomes for All" by George Church, in the January 2006 issue of Scientific American. To learn more about challenges related to visualizing genomes, see "Similarities Between Human and Chimp Genomes Revealed by Hilbert Curve" by Martin Krzywinski).
.png?w=510)
Graphic by Mark Kunitomi
Each of the colored lines in Kunitomi's graphic represents a string of chemical base pairs—the A,T, C and G of the mosquito's genetic code—whose accuracy researchers are highly confident about. These precisely known chemical base pair sequences are known as contigs. The detail below shows six of them.

There are 3,752 contigs in the full map. The 2007 draft map included 36,206 contigs. The ultimate goal of continued sequencing efforts is to end up with just three lines; one continuous string of base pairs for each chromosome.
The length of each colored line represents the number of base pairs in a contig, ranging from about 35,000 (smallest visible line on the graphic) to 7,901,702. The full data set of this cell line of A. aegypti is comprised of about 1.7 billion base pairs, which includes both coding regions (genes) and non-coding regions of the genome.
Each grouping of colored lines represents contigs that the researchers are pretty sure belong together, but some gaps, overlaps, conflicts, and/or other uncertainties may exist at the points of connection (circled in black, below).
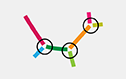
The position of each group within the full image grid is roughly based on size. Line shape (curves, squiggles, and loops) and orientation are arbitrary.
Kunitomi created the graphic with the bioinformatics visualization tool Bandage, developed by Ryan Wick (currently a research assistant in Kathryn Holt's research group at University of Melbourne). A description paper was published last year in the journal Bioinformatics: the software is available online, or you can clone the source code on GitHub.
The bottom line? Researchers have made significant steps toward piecing together the genome of Aedes aegypti, but the map is still quite fragmented. Visualizations like this one allow researchers to zoom in and identify which regions still need more work, and allow non-specialists—like me—to track their progress.