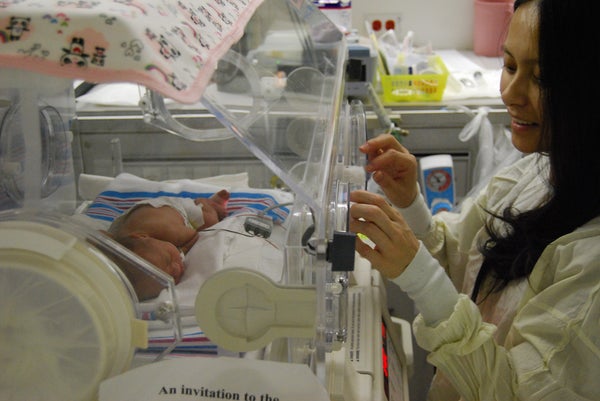
This article was published in Scientific American’s former blog network and reflects the views of the author, not necessarily those of Scientific American
If you have a google news alert set for “Antibiotic Resistance” (or just read Maryn McKenna) like I do, you can’t help but get scared. In the war against infectious bacteria, we’re on the retreat. From the economic barriers to antibiotic discovery to the appearance of a new superbug in the US, it increasingly seems that all of the salvos in this war are being launched by the enemy.
Perhaps it’s not surprising then that not even infants are spared from combat. Research published earlier this year analyzed the development of the gut microbiome of preterm infants and suggests that the risks go beyond low birthweight.
Developmental dynamics of the preterm infant gut microbiota and antibiotic resistome
On supporting science journalism
If you're enjoying this article, consider supporting our award-winning journalism by subscribing. By purchasing a subscription you are helping to ensure the future of impactful stories about the discoveries and ideas shaping our world today.
All of the infants in this study were given at least some antibiotics shortly after birth. This isn’t surprising - preterm infants are at substantially higher risk for infection. Then again, it’s also not surprising that these infants had significantly altered microbial populations in their guts. Previous work suggests that having a lot of diversity (lots of different species) in the gut is a sign of a healthy microbiome, but preterm infants had substantially lower diversity than babies that had gone to term.
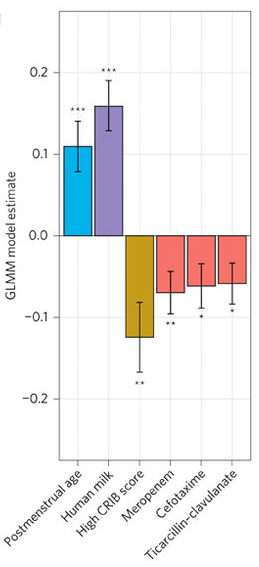
Caption
Treatment with antibiotics does not cull all microbes to the same extent, so this result is not unexpected, but it’s clear that certain antibiotics had more of an impact than others. Interestingly, the farther along these babies were before they were born (postmenstrual age) the more diverse their microbes, and feeding with human breast milk rather than formula also improved diversity. It’s unclear if these factors are causally linked to antibiotic use, though babies born earlier are typically less healthy and more likely to receive longer courses of antibiotics.
These data are clearly of interest to Physicians that deal with preterm infants, but the news for the rest of us starts in Figure 2. The authors reasoned that this population of tiny people that spend a large amount of time in a hospital and are treated early with antibiotics would be likely to harbor antibiotic-resistant bacteria. To test this hypothesis, they gathered metagenomic data to look for the presence of potential antibiotic resistance genes.
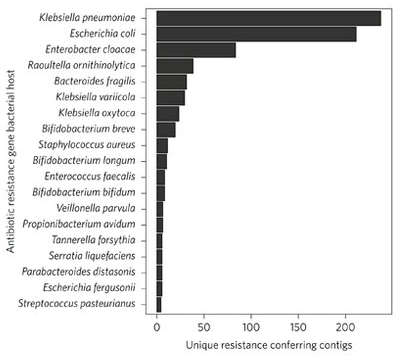
Caption
The three species (Escherichia coli, Enterobacter cloacae and Klebsiella pneumoniae) encoding the highest number of AR [antibiotic resistance] genes include organisms belonging to the ESKAPE pathogen family and are widely distributed in NICU environments
Many of these resistance genes have not been characterized but are expected to lead to resistance because they look like other resistance genes. Even more distressing than the shear number of AR genes they uncovered is the fact that enrichment for resistance was not restricted to the type of antibiotic used. There are many different classes of antibiotics that target different parts of the bacterial life cycle. Often, genes that allow microbes to resist one type of antibiotic will also increase resistance to antibiotics in the same class. At the same time, you would only expect antibiotic treatment to select for resistance to that class. But these authors found that treating these infants with one class of antibiotic led to increased abundance of genes for resistance to many other classes.
There are manyideas being offered to combat AR, but the discovery of a new front that I didn’t even consider we were fighting on is still a bit depressing.