This article was published in Scientific American’s former blog network and reflects the views of the author, not necessarily those of Scientific American
The microbiome is in the news again in a big way. Last week, the Obama Administration announced a major new funding initiative geared at spurring research into our microbial ecosystems, prompting tons of mediacoverage. More money for research is great, though I have some concerns about the ways directed spending might distort scientific progress (eg. $500 million towards science generally could also be very well spent).
But getting a handle on the gut microbiome is hard. Basically everything can affect it or be affected by it. Some of the things are obvious, like diet or antibiotic use or when you last pooped, but others aren’t as obvious, like time of day, background genetics, autoimmune disease in the brain and infectious diseases. Many studies have reported differences in different patient cohorts, but teasing out which of these differences are actually meaningful, or in which direction the causal arrow points is even harder (eg. does MS cause changes to the microbiome, or do different microbiomes cause MS, or does something unknown alter both the microbiome and prevalence of MS?)
A lot of this uncertainty comes from a fundamental lack of data. The differences in the microbiome between two healthy people can be so dramatic that potential differences between healthy and unhealthy people can get lost in the noise. A few years ago - there was good evidence that a particular microbe was strongly associated with a healthy gut community, until a population of humans that completely lack that microbe (with no apparent ill effects) were discovered. That population, the Hazda of Tanzania, are a group of hunter-gatherers, and their guts were wildly different than those of the (largely) American and western Europeans that had been studied before.
On supporting science journalism
If you're enjoying this article, consider supporting our award-winning journalism by subscribing. By purchasing a subscription you are helping to ensure the future of impactful stories about the discoveries and ideas shaping our world today.
But knowing that they’re different doesn’t really help us in understanding why they’re different. Is it because their diet is different, is it their lifestyle, or just the fact that they’re on a different continent? A study published earlier this year attempts to edge us closer to an answer:
This study analyzed the gut microbiomes of two different populations of people - both are indigenous to the same region of central Africa, but one group (the Bantu) have an agrarian society and have what the authors call a “Western-like” diet (though I have a sneaking suspicion that I wouldn’t recognize much of the cuisine), and the BaAka are rain forest hunter-gatherers.
The most important (or at least most readily accessible) bit of analysis I think is in Figure 3 where the authors look at the “UniFrac Distance” between these two groups of indigenous people and a cohort from the US. This measure is a way of breaking down complex data on thousands of microbial species into a single measure. Unweighted UniFrac looks only at what species are present, while the weighted measure also takes into account how much of each bug are there.
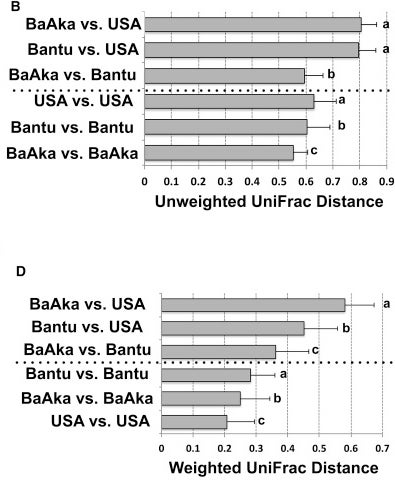
From doi:10.1016/j.celrep.2016.02.013 CC-BY
There are differences between all the populations to be sure, but it’s striking how much variation there is within the populations as well (the bars below the dotted lines are comparing within groups). Another way to look at the same data is to perform a principal components analysis, which again is a mathematical way of ordering complexity - you can watch that video to get a sense of how it’s done, but the short version is that we can peel out the variables that explain the majority of the variation.
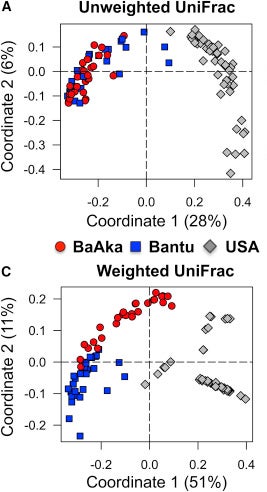
From doi:10.1016/j.celrep.2016.02.013 CC-BY
Here, each dot is a single person whose microbiome was sampled, and the axes represent those principal components. What you can see is that when we don’t account for how much of each species is present, the two indigenous populations are almost identical, though a couple of the Bantu folks look a bit more like the Americans. This could be due to location, or because the diets of the indigenous folks are more similar than we thought, or maybe it’s because they get malaria at similar rates. Who knows? When they do account for abundances, the populations split a bit more, but here it seems to be the BaAka that look more like the Americans (at least along PC1, which supposedly accounts for over half of the variation they measure).
Both indigenous groups have markedly more diverse microbial compositions than the American group, and some previous studies have shown that lower microbial diversity is linked to poor health outcomes, but I’m not sure we can read much into that. These groups are so diverse, both within cohorts and between cohorts, I’m skeptical that the variation that’s seen here can tell us much. I would have like to see a comparison to a more westernized or urban population in Africa as the comparison group rather than people from the US, but I’m not sure even that would be informative.
Researchers could probably suck up all $500 million that the Obama administration has allocated just in sampling different populations in different locations and at different times of day and with different health conditions, and we might be no closer to getting a handle on the staggering complexity that is our gut ecosystem. But we’ll keep inching towards more nuance, and hopefully someday we can put this understanding to good use.